The Genetics of Glaucoma
All content on Eyewiki is protected by copyright law and the Terms of Service. This content may not be reproduced, copied, or put into any artificial intelligence program, including large language and generative AI models, without permission from the Academy.
Description/Overview
Glaucoma is a heterogeneous group of optic neuropathies characterized by progressive retinal ganglion cell loss and corresponding visual field damage. Although elevated intraocular pressure (IOP) remains the most important modifiable risk factor, genetic influences play a major role in disease susceptibility, age at onset, phenotype, and rate of progression. Glaucoma genetics spans a spectrum from rare, high-penetrance Mendelian variants causing early-onset disease to common low-effect variants that contribute to the risk of adult-onset glaucoma in a polygenic manner. This genetic framework is increasingly important for diagnosis, family counseling, risk stratification, and the future development of targeted therapies.[1][2][3][4]
Genetic Architecture of Glaucoma
The genetic basis of glaucoma can broadly be divided into monogenic and complex polygenic disease. Monogenic forms are most relevant in primary congenital glaucoma (PCG), juvenile open-angle glaucoma (JOAG), and some familial normal-tension glaucoma (NTG). In these settings, a pathogenic variant may have a relatively large effect size and can sometimes explain the phenotype in a single family. By contrast, the more common adult-onset forms of primary open-angle glaucoma (POAG) and primary angle-closure glaucoma (PACG) usually reflect the cumulative effect of many susceptibility loci together with environmental and systemic influences.[1][3]
Large genome-wide association studies have substantially expanded the known genetic landscape of POAG. A major cross-ancestry meta-analysis identified 127 loci associated with open-angle glaucoma, highlighting pathways related to IOP regulation, extracellular matrix biology, vascular regulation, and optic nerve vulnerability. These studies support the concept that glaucoma is not a single-gene disorder in most adults, but rather a biologically diverse disease with multiple contributing pathways.[3]
Key Genes and Genotype–Phenotype Correlations
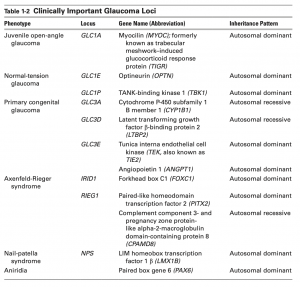
Several genes are especially important in clinical glaucoma genetics. MYOC is the best-established gene in juvenile open-angle glaucoma and is typically inherited in an autosomal dominant pattern. MYOC-associated disease often presents with markedly elevated IOP at a young age and may be more severe than typical adult-onset POAG. Because of its relatively strong genotype-phenotype correlation, MYOC is one of the most clinically useful genes in targeted testing for early-onset open-angle glaucoma.[1][2][3][4]
In primary congenital glaucoma, CYP1B1 remains the most important gene worldwide, particularly in populations with consanguinity or founder effects. Other genes implicated in childhood glaucoma and developmental outflow abnormalities include LTBP2, TEK, and ANGPT1. TEK is especially notable because it causes an autosomal dominant form of congenital glaucoma with variable expressivity, emphasizing that even within Mendelian disease, penetrance may be incomplete and clinical severity may differ substantially between affected relatives.[1][2][3][5][4]
For normal-tension glaucoma, OPTN and copy number variation involving TBK1 are the most classically recognized genetic contributors. TBK1-associated disease may present with significant optic nerve cupping and progression despite statistically normal IOP, making it especially relevant in patients with strong family history, early disease, or unexpectedly severe NTG. More recently, METTL23 has emerged as another potentially important gene in familial NTG, though its exact place in routine clinical testing is still evolving. [1][2][6][4]
Syndromic and anterior segment dysgenesis–associated glaucomas also have important genetic causes. FOXC1 and PITX2 are strongly associated with Axenfeld-Rieger spectrum and can help explain glaucoma occurring in association with iris, angle, or corneal developmental abnormalities. Recognition of these genes is useful not only for the proband but also for systemic evaluation and family counseling.[1][2][4]
Polygenic Risk and Common Adult-Onset Glaucoma
Most adult-onset POAG is genetically complex. Common variants in loci such as CDKN2B-AS1, SIX6, and TMCO1 do not establish a diagnosis on their own, but they contribute to disease susceptibility and are increasingly used in polygenic risk score (PRS) models. The clinical value of PRS is not to replace examination-based diagnosis, but rather to refine risk assessment in selected populations.[3][7]
Recent work has strengthened the case for PRS in glaucoma. In a 2024 post hoc analysis of the Ocular Hypertension Treatment Study, a POAG polygenic risk score helped distinguish ocular hypertensive individuals at lower versus higher long-term risk of conversion to glaucoma. This suggests that PRS may eventually help guide surveillance intensity or identify patients most appropriate for early intervention. However, an important limitation is that PRS performance varies by ancestry, and currently available models have generally performed better in European-ancestry cohorts than in African-ancestry populations. This remains a major barrier to equitable implementation.[8][9]
The genetic basis of primary angle-closure disease is also increasingly recognized as polygenic. A 2024 systematic review and meta-analysis confirmed multiple associated loci and highlighted genes involved in ocular biometry and extracellular matrix regulation, including ATOH7, CALCRL, FBN1, LOXL1, MMP19, and VAV3. Despite this progress, the clinical application of these findings remains limited, particularly for predicting which angle-closure suspects will progress.[10]
Clinical Role of Genetic Testing
Genetic testing is most useful when the phenotype suggests a high likelihood of Mendelian disease. This includes congenital or childhood-onset glaucoma, juvenile-onset open-angle glaucoma, glaucoma associated with anterior segment dysgenesis or syndromic features, and familial normal-tension glaucoma. In these scenarios, testing may confirm a diagnosis, clarify inheritance, guide screening of relatives, and occasionally influence management intensity.[2][11]
By contrast, routine diagnostic genetic testing is not currently recommended for typical late-onset POAG without unusual features, because most cases are genetically complex and currently identified risk variants do not have sufficient specificity for stand-alone clinical decision-making. The American Academy of Ophthalmology has specifically advised caution in using genetic testing for genetically complex eye diseases until results are clearly actionable and appropriately interpreted. Variant interpretation should follow accepted clinical genetics standards, including ACMG/AMP terminology such as pathogenic, likely pathogenic, and variant of uncertain significance. [2][11]
Counseling and Family Screening
The most immediate clinical benefit of glaucoma genetics is often family screening. Identification of a pathogenic variant in a proband can justify earlier and more targeted ophthalmic surveillance in at-risk relatives, including IOP measurement, gonioscopy when appropriate, and optic nerve and visual field assessment. Counseling is essential because many glaucoma-associated variants show incomplete penetrance and variable expressivity, and a negative test does not eliminate risk in a strongly affected family. Pre-test and post-test counseling should therefore address diagnostic yield, uncertainty, psychosocial implications, and the limitations of current knowledge.[2][11]
Emerging Directions
Glaucoma genetics is increasingly moving from risk discovery toward translational application. Current research is integrating GWAS data with single-cell transcriptomics and regulatory biology to identify disease-relevant pathways and potential therapeutic targets. At the same time, gene-based therapies are beginning to enter early clinical development. Recent reviews describe growing interest in gene transfer and gene editing strategies aimed at improving aqueous outflow, reducing trabecular meshwork dysfunction, or protecting retinal ganglion cells. Although these approaches remain investigational, they represent an important future direction for the field.[12][13]
Conclusion
Glaucoma genetics is now central to understanding disease heterogeneity. Rare Mendelian variants are most clinically relevant in congenital, juvenile, syndromic, and familial NTG phenotypes, while common variants contribute to the inherited burden of adult-onset glaucoma in a polygenic manner. At present, the greatest clinical utility lies in phenotype-directed genetic testing and cascade screening of families at risk. In the future, improved PRS models, broader ancestry representation, and gene-targeted therapies may further integrate genetics into routine glaucoma care.
References
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 Roy S. Emerging strategies targeting genes and cells in glaucoma. Vision Res. 2025;227:108533. doi:10.1016/j.visres.2024.108533
- ↑ 2.0 2.1 2.2 2.3 2.4 2.5 2.6 2.7 Gedde SJ, Vinod K, Wright MM, et al. Primary Open-Angle Glaucoma Preferred Practice Pattern®. Ophthalmology. 2021;128(1):P71-P150. doi:10.1016/j.ophtha.2020.10.022
- ↑ 3.0 3.1 3.2 3.3 3.4 3.5 Gharahkhani P, Jorgenson E, Hysi P, et al. Genome-wide meta-analysis identifies 127 open-angle glaucoma loci with consistent effect across ancestries. Nat Commun. 2021;12(1):1258. Published 2021 Feb 24. doi:10.1038/s41467-020-20851-4
- ↑ 4.0 4.1 4.2 4.3 4.4 American Academy of Ophthalmology. Basic and Clinical Science Course (BCSC): Section 10 – Glaucoma. 2024–2025 ed. San Francisco, CA: American Academy of Ophthalmology; 2024.
- ↑ Souma T, Tompson SW, Thomson BR, et al. Angiopoietin receptor TEK mutations underlie primary congenital glaucoma with variable expressivity. J Clin Invest. 2016;126(7):2575-2587. doi:10.1172/JCI85830
- ↑ Scheetz TE, Tollefson MR, Roos BR, et al. METTL23 Variants and Patients With Normal-Tension Glaucoma. JAMA Ophthalmol. 2024;142(11):1037-1045. doi:10.1001/jamaophthalmol.2024.3829
- ↑ He W, Lee SS, Diaz Torres S, et al. Predictive Power of Polygenic Risk Scores for Intraocular Pressure or Vertical Cup-Disc Ratio. JAMA Ophthalmol. 2025;143(1):15-22. doi:10.1001/jamaophthalmol.2024.4856
- ↑ Sekimitsu S, Ghazal N, Aziz K, et al. Primary Open-Angle Glaucoma Polygenic Risk Score and Risk of Disease Onset: A Post Hoc Analysis of a Randomized Clinical Trial. JAMA Ophthalmol. Published online November 7, 2024. doi:10.1001/jamaophthalmol.2024.4376
- ↑ Chang-Wolf JM, Kinzy TG, Driessen SJ, et al. Performance of Polygenic Risk Scores for Primary Open-Angle Glaucoma in Populations of African Descent. JAMA Ophthalmol. 2025;143(1):7-14. doi:10.1001/jamaophthalmol.2024.4784
- ↑ Liang YJ, Wang YY, Rong SS, et al. Genetic Associations of Primary Angle-Closure Disease: A Systematic Review and Meta-Analysis. JAMA Ophthalmol. 2024;142(5):437-444. doi:10.1001/jamaophthalmol.2024.0363
- ↑ 11.0 11.1 11.2 Stone EM, Aldave AJ, Drack AV, et al. Recommendations for genetic testing of inherited eye diseases: report of the American Academy of Ophthalmology task force on genetic testing. Ophthalmology. 2012;119(11):2408-2410. doi:10.1016/j.ophtha.2012.05.047
- ↑ Anton N, Geamănu A, Iancu R, et al. A Mini-Review on Gene Therapy in Glaucoma and Future Directions. Int J Mol Sci. 2024;25(20):11019. Published 2024 Oct 14. doi:10.3390/ijms252011019
- ↑ Hamel AR, Yan W, Rouhana JM, et al. Integrating genetic regulation and single-cell expression with GWAS prioritizes causal genes and cell types for glaucoma. Nat Commun. 2024;15(1):396. Published 2024 Jan 9. doi:10.1038/s41467-023-44380-y